Using the computing powers of the Texas Advanced Computing Center, TACC, at UT-Austin, researchers are making progress on the biological mechanisms of West Nile virus. The computational heavy lifting was done by the TACC’s Stampede2, Jetstream and Lonestar 5.
“Humans get infected with West Nile virus by the bite of an infected mosquito,” said Margo Brinton, lead author and professor of biology at Georgia State University. “The virus replicates primarily in white blood cells in the blood. In a few people, it can cross the blood-brain barrier and replicate in the brain’s neurons, causing (inflammation of the brain).”
In present day, the virus is now in Africa, Southern Asia, Australia, the Middle East, North and Central America and Europe, according to Brinton.
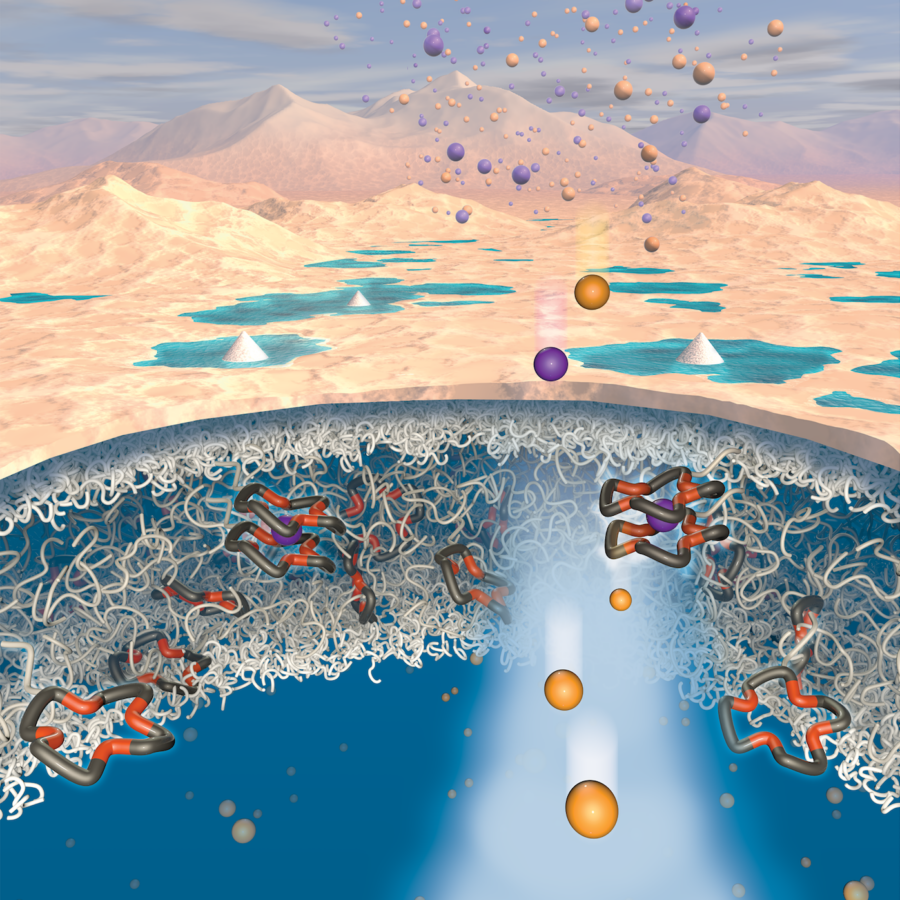
West Nile virus has an RNA genome, or genetic material, which it can replicate in the cells it infects. Certain conserved structures, including a stem loop on the RNA strand, have shown to play an important role in regulating the ability of the virus to replicate its genome.
The researchers had previously discovered a protein called TIAR that interacts with this stem loop structure in RNA. Now, in a paper published in Analytical Chemistry from January 2017, they found that the specific ratio of TIAR proteins to the stem loop in RNA is 4-to-1.
The researchers said it was important because they realized how much more complex the viral interactions in the cell can be with RNA than previously thought.
“We think the cellular protein TIAR acts to enhance viral genome production inside infected cells,” Brinton said. “We are continuing to define this complex viral RNA-cell protein interaction by using other physical chemistry and biochemical techniques.”
Fellow author on the paper and University of Texas Health Science Center researcher Borries Demeler works with a technique called analytical ultracentrifugation, which played an important role in studying West Nile virus.
“Analytical ultracentrifugation is a separation technique in which high centrifugal forces are used to separate molecules of different sizes and shapes mixed in a solution,” Demeler said. “It can then measure the size and shape of different kinds of molecules and indicate if they are interacting, and if so, how strongly.”
Demeler developed analysis software for research that uses analytical ultracentrifugation; however, information processing can require more significant computational power.
“The separation methods used in (analytical ultracentrifugation) are computationally extremely complex and require a demanding algorithm for modeling,” Demeler said. “Due to the very large data amounts, large computers are needed to extract this information, which was done on supercomputers at the TACC.”
Demeler also said researchers used a new technology, a multiwavelength detector, which can separate molecules based on their optical properties in addition to size and shape. Because different types of molecules absorb light at different wavelengths, biological molecules like DNA, RNA, carbohydrates and lipids all show different absorbance patterns.
Using the high sensitivity of the detector, the researchers were able to separate their biological samples.
“It gives us a new way to look at complex interactions occurring in a living cell,” Demeler said.
Using these same computational techniques, Demeler said he expects a wide variety of applications in biological research, especially by studying the interaction between two dissimilar biological molecules like DNA, RNA or proteins.
“The problem we tackled with the West Nile virus is just one example of a nearly infinite range of interactions and systems that could be studied using this technique,” Demeler said. “I expect many new discoveries will result from this ability to examine mixtures in greater detail and help answer important questions in biochemical research, contributing to developing cures for cancer and many other diseases.”